Bacterial biofilms are dense, surface-attached communities composed of microorganisms that exhibit multicellular behavior. These structures play a major role in medicine, the environment, and industry by resisting antibiotics, surviving cleaning agents, and protecting microbial populations under stress. Despite decades of study, much remains unknown about how biofilm architecture supports this resilience—especially the role of rigid 3D structures formed within the biofilm matrix.
This project investigates how the spatial organization and physical properties of biofilms influence bacterial stress responses, survival, and growth. Using Enterococcus faecalis and Escherichia coli as model organisms, we explore how rigid structures arise through two distinct mechanisms: aggregation of bacterial cells and the production of amyloid fibers. Our findings suggest that tightly packed clusters formed by cell aggregation alone can generate heterologous rigid structures—without the need for specialized matrix components—creating sheltered microenvironments that allow bacteria to survive even high doses of antibiotics.
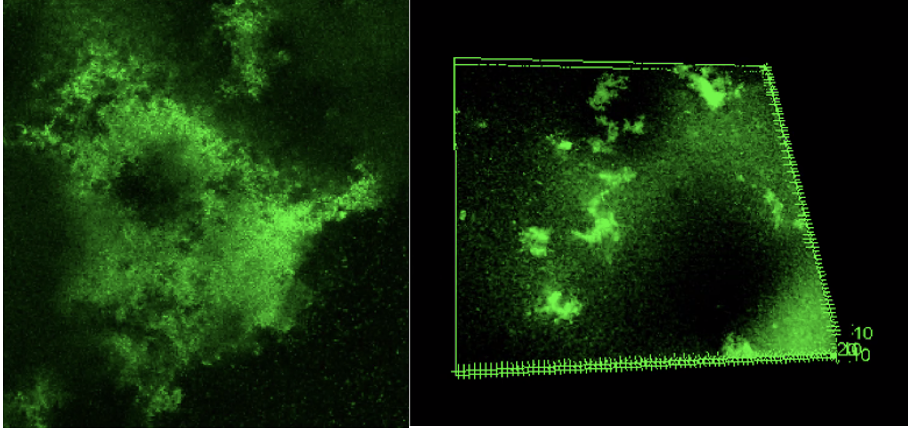
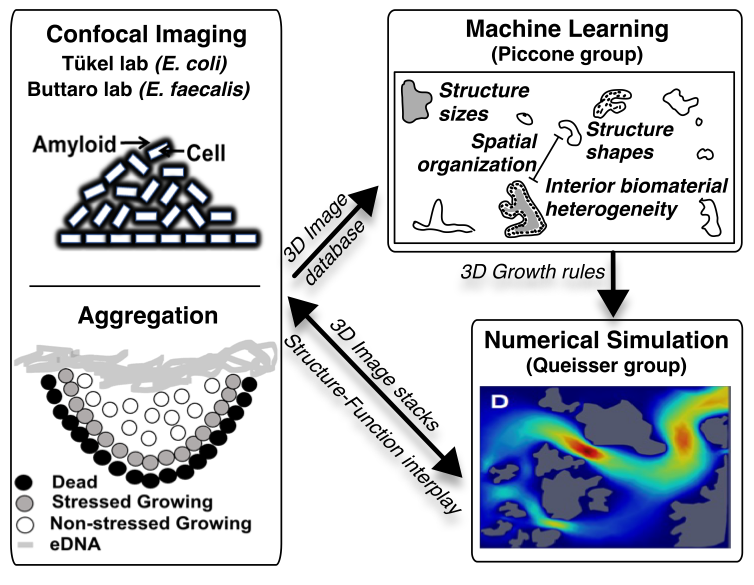
Through a multidisciplinary approach that integrates biological experiments, machine learning, and computational modeling, we aim to uncover the spatial rules and growth patterns that govern the formation and function of these rigid structures. High-resolution 3D confocal imaging is used to capture biofilm structure under both static and flow conditions, while fluorescent probes reveal gene expression linked to stress and structure formation. Machine learning models analyze thousands of images to detect patterns in size, density, and spatial arrangement, identifying whether these characteristics are species-specific or governed by general rules. Meanwhile, simulation tools model how antibiotics and other stress signals move through the biofilm, using diffusion-advection-reaction frameworks to predict protected zones within the microbial community.
Our goal is to understand how stress signaling is redistributed through 3D architectures in biofilms and how this leads to the emergence of non-stressed, viscous microenvironments that support continued growth. These findings will have broad applications—from disrupting harmful biofilms in clinical settings to understanding microbial persistence in environmental or industrial systems. The research also lays the groundwork for studying mixed-species biofilms, where some organisms may not form rigid structures but still benefit from the protection offered by neighboring species that do.
In parallel with our scientific aims, we are committed to open science. All biofilm images, computational models, and machine learning tools will be made publicly available through an open-access repository, hosted by the Neural Engineering Data Consortium (NEDC). This platform will support a wide range of users, including microbiologists, mathematicians, engineers, educators, and students. It will also serve as an educational resource, offering annotated datasets and training opportunities for high school, undergraduate, and graduate students.
Ultimately, this project brings together biology, data science, and engineering to answer fundamental questions about how bacterial communities survive under pressure. By revealing the rules of biofilm architecture and stress adaptation, we hope to develop strategies that can either protect beneficial microbial communities—such as those in the gut—or break down harmful biofilms in medicine, agriculture, and industry.